Deep Learning and Feature Extraction of Brain Tumour Detection
Keywords:
Brain tumor, Image acquisition, Deep learning algorithm, MRI ImagingAbstract
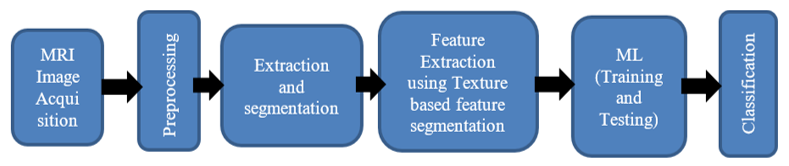
In medical imaging, automated flaw detection has grown in importance. The ability to forecast tumor (brain) detection on one's own during an MRI scan is essential for preparing patients. Conventional methods of calculating z are developed to facilitate the work of radiologists. The size and variety of molecular structures in brain tumours presents a challenge for MRI diagnosis. This research uses deep learning (DL) techniques including support vector machines (SVM), artificial neural networks (ANN), and convolutional ANN to detect tumours in MRI scans (CNN). Segmentation scanning, feature extraction, and brain tumor classification are steps in the recommended technique. The second step consists of dataset preparation and input picture scanning. The third step involves figuring out how to extract features from a scanned picture. A number of machine learning methods are then utilized to classify the data according on these criteria. One of the most well-known neural networks (CNN) is employed in this article to differentiate between different kinds of MRI tumors.
Downloads
References
M. S. Goswami and M. L. K. P. Bhaiya, “Brain tumour detection using unsupervised learning based neural network,” in Proceedings - 2013 International Conference on Communication Systems and Network Technologies, CSNT 2013, Apr. 2013, pp. 573–577. doi: 10.1109/CSNT.2013.123.
T. Rajesh and R. S. M. Malar, “Rough set theory and feed forward neural network based brain tumor detection in magnetic resonance images,” in International Conference on Advanced Nanomaterials & Emerging Engineering Technologies, Jul. 2013, pp. 240–244. doi: 10.1109/ICANMEET.2013.6609287.
S. Rajeshwari and T. S. Sharmila, “Efficient quality analysis of MRI image using preprocessing techniques,” in 2013 IEEE Conference on Information and Communication Technologies, ICT 2013, Apr. 2013, pp. 391–396. doi: 10.1109/CICT.2013.6558127.
T. A. Assegie, R. L. Tulasi, and N. K. Kumar, “Breast cancer prediction model with decision tree and adaptive boosting,” IAES International Journal of Artificial Intelligence (IJ-AI), vol. 10, no. 1, pp. 184–190, Mar. 2021, doi: 10.11591/ijai.v10.i1.pp184-190.
D. Singh and K. Kaur, “Classification of abnormalities in brain MRI images using GLCM , PCA and SVM,” International Journal of Engineering and Advanced Technology (IJEAT), vol. 1, no. 6, pp. 243–248, 2012.
P. Gadpayleand and P. S. Mahajani, “Detection and classification of brain tumor in MRI images,” International Journal of Emerging Trends in Electrical and Electronics (IJETEE), vol. 5, no. 1, 2013.
M. Shasidhar, V. S. Raja, and B. V. Kumar, “MRI brain image segmentation using modified fuzzy c-means clustering algorithm,” in Proceedings - 2011 International Conference on Communication Systems and Network Technologies, CSNT 2011, Jun. 2011, pp. 473–478. doi: 10.1109/CSNT.2011.102.
K. Sharma and N. Kaur, “Comparative analysis of various edge detection techniques,” International Journal of Advanced Research in Computer Science and Software Engineering, vol. 3, no. 12, pp. 617 – 621, 2013.
J. Selvakumar, A. Lakshmi, and T. Arivoli, “Brain tumor segmentation and its area calculation in brain MR images using K-mean clustering and Fuzzy C-mean algorithm,” in IEEE-International Conference on Advances in Engineering, Science and Management, ICAESM-2012, 2012, pp. 186–190. [Online]. Available: https://ieeexplore.ieee.org/abstract/document/6215996/ metrics#metrics
R. J. Ramteke and K. M. Yashawant, “Automatic medical image classification and abnormality detection using k-nearest neighbour,” International Journal of Advanced Computer Research, vol. 2, no. 4, pp. 190–196, 2012, [Online]. Available: https://www.researchgate.net/publication/305403850_Automatic_Medical_Image_Classification_and_Abnormality_Detection_Using_K-_Nearest_Neighbour
X. Xuan and Q. Liao, “Statistical structure analysis in MRI brain tumor segmentation,” in Proceedings of the 4th International Conference on Image and Graphics, ICIG 2007, Aug. 2007, pp. 421–426. doi: 10.1109/ICIG.2007.181.
K. Sharma, A. Kaur, and S. Gujral, “Brain tumor detection based on machine learning algorithms,” International Journal of Computer Applications, vol. 103, no. 1, pp. 7–11, Oct. 2014, doi: 10.5120/18036-6883.
N. B. Bahadure, A. K. Ray, and H. P. Thethi, “Image analysis for MRI based brain tumor detection and feature extraction using biologically inspired BWT and SVM,” International Journal of Biomedical Imaging, pp. 1–12, 2017, doi: 10.1155/2017/9749108.
M. W. Nadeem et al., “Brain tumor analysis empowered with deep learning: A review, taxonomy, and future challenges,” Brain Sciences, vol. 10, no. 2, pp. 1–33, Feb. 2020, doi: 10.3390/brainsci10020118.
D. V. Gore and V. Deshpande, “Comparative study of various techniques using deep learning for brain tumor detection,” in 2020 International Conference for Emerging Technology (INCET), Jun. 2020, pp. 1–4. doi: 10.1109/INCET49848.2020.9154030.
W. Widhiarso, Y. Yohannes, and C. Prakarsah, “Brain tumor classification using gray level co-occurrence matrix and convolutional neural network,” IJEIS (Indonesian Journal of Electronics and Instrumentation Systems), vol. 8, no. 2, pp. 179–190, Oct. 2018, doi: 10.22146/ijeis.34713.
H. Mohsen, E.-S. A. El-Dahshan, E.-S. M. El-Horbaty, and A.-B. M. Salem, “Classification using deep learning neural networks for brain tumors,” Future Computing and Informatics Journal, vol. 3, no. 1, pp. 68–71, Jun. 2018, doi: 10.1016/j.fcij.2017.12.001.
K. Kamnitsas et al., “Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation,” Medical Image Analysis, vol. 36, pp. 61–78, Feb. 2017, doi: 10.1016/j.media.2016.10.004.
S. K. Chandra and M. K. Bajpai, “Effective algorithm for benign brain tumor detection using fractional calculus,” in TENCON 2018 - 2018 IEEE Region 10 Conference, Oct. 2018, pp. 2408–2413. doi: 10.1109/TENCON.2018.8650163.
S. W. Kareem, R. Z. Yousif, and S. M. J. Abdalwahid, “An approach for enhancing data confidentiality in hadoop,” Indonesian Journal of Electrical Engineering and Computer Science, vol. 20, no. 3, pp. 1547–1555, Dec. 2020, doi: 10.11591/ijeecs.v20.i3.pp1547-1555.
A. Triayudi and I. Fitri, “Comparison of the feature selection algorithm in educational data mining,” TELKOMNIKA (Telecommunication Computing Electronics and Control), vol. 19, no. 6, pp. 1865–1871, Dec. 2021, doi: 10.12928/telkomnika.v19i6.21594.
J. Seetha and S. S. Raja, “Brain tumor classification using Convolutional Neural Networks,” Biomedical and Pharmacology Journal, vol. 11, no. 3, pp. 1457–1461, Sep. 2018, doi: 10.13005/bpj/1511.
J. Cheng et al., “Enhanced performance of brain tumor classification via tumor region augmentation and partition,” PLoS ONE, vol. 10, no. 10, pp. 1–13, Oct. 2015, doi: 10.1371/journal.pone.0140381.
G. S. Tandel et al., “A review on a deep learning perspective in brain cancer classification,” Cancers, vol. 11, no. 1, pp. 1–32, Jan. 2019, doi: 10.3390/cancers11010111.
G. Litjens et al., “A survey on deep learning in medical image analysis,” Medical Image Analysis, vol. 42, pp. 60–88, Dec. 2017, doi: 10.1016/j.media.2017.07.005.
L. Szilagyi, L. Lefkovits, and B. Benyo, “Automatic brain tumor segmentation in multispectral MRI volumes using a fuzzy c-means cascade algorithm,” in 2015 12th International Conference on Fuzzy Systems and Knowledge Discovery (FSKD), Aug. 2015, pp. 285–291. doi: 10.1109/FSKD.2015.7381955.
M. Soltaninejad et al., “Automated brain tumour detection and segmentation using superpixel-based extremely randomized trees in FLAIR MRI,” International Journal of Computer Assisted Radiology and Surgery, vol. 12, no. 2, pp. 183–203, Feb. 2017, doi: 10.1007/s11548-016-1483-3.
J. R. Fink, M. Muzi, M. Peck, and K. A. Krohn, “Continuing education: Multi-modality brain tumor imaging – MRI, PET, and PET/MRI,” Journal of Nuclear Medicine, vol. 56, no. 10, pp. 1554–1561, Oct. 2015, doi: 10.2967/jnumed.113.131516.
G. M. Shivakumarswamy, P. V Akshay, T. A. Chethan, B. H. Prajwal, and S. V Hande, “Brain tumour detection using Image processing and sending tumour information over GSM,” International Journal of Advanced Research in Computer and Communication Engineering, vol. 5, no. 5, pp. 179–183, 2016, doi: 10.17148/IJARCCE.2016.5543.
Y. Jia, Y. Zhang, and T. Rabczuk, “A novel dynamic multilevel technique for image registration,” Computers & Mathematics with Applications, vol. 69, no. 9, pp. 909–925, May 2015, doi: 10.1016/j.camwa.2015.02.010.
T. Kalaiselvi, P. Nagaraja, and P. Sriramakrishnan, “A simple image processing approach to abnormal slices detection from MRI tumor volumes,” The International journal of Multimedia & Its Applications, vol. 8, no. 1, pp. 55–64, Feb. 2016, doi: 10.5121/ijma.2016.8105.
R. Kumar and K. J. Mathai, “Brain tumor segmentation by modified k-mean with morphological operations,” International Journal of Innovative Research in Science, Engineering and Technology, vol. 6, no. 8, pp. 16191–16198, 2017, doi: 10.15680/IJIRSET.2016.0608079.
A. Mukaram, C. Murthy, and M. Z. Kurian, “An automatic brain tumour detection, segmentation and classification using MRI image,” International Journal of Electronics, Electrical and Computational System, vol. 6, no. 5, pp. 54–65, 2017, [Online]. Available: https://www.semanticscholar.org/paper/An-Automatic-Brain-Tumour-Detection%2C-Segmentation-Mukaram-Murthy/92e91674497afffd2acca94d42ac39513e10e184
S. Patel and D. Rao, “Brain tumor detection in MRI images with new multiple thresholding,” Journal of Network Communications and Emerging Technologies (JNCET), vol. 7, no. 6, pp. 34–39, 2017, [Online]. Available: https://www.jncet.org/Manuscripts/Volume-7/Issue-6/Vol-7-issue-6-M-08.pdf
A. Pawar et al., “Adaptive FEM-based nonrigid image registration using truncated hierarchical B-splines,” Computers & Mathematics with Applications, vol. 72, no. 8, pp. 2028–2040, Oct. 2016, doi: 10.1016/j.camwa.2016.05.020.
R. Sharmila and K. S. Joseph, “Brain tumour detection of MR image using Naïve Beyer classifier and support vector machine,” nternational Journal of Scientific Research in Computer Science, Engineering and Information Technology, vol. 3, no. 3, pp. 690–695, 2018, [Online]. Available: https://1library.net/document/zp2wwroy-brain-tumour-detection-naive-classifier-support-vector-machin.html
M. Havaei, F. Dutil, C. Pal, H. Larochelle, and P.-M. Jodoin, “A convolutional neural network approach to brain lesion segmentation,” in Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries, Chsm: Springer International Publishing, 2016, pp. 195–208. doi: 10.1007/978-3-319-30858-6_17.
S. Pereira, A. Pinto, V. Alves, and C. A. Silva, “Deep convolutional neural networks for the segmentation of gliomas in multi-sequence MRI,” in Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries, Chsm: Springer International Publishing, 2016, pp. 131–143. doi: 10.1007/978-3-319-30858-6_12.
Kamnitsas, K.; Ledig, C.; Newcombe, V.F.; Simpson, J.P.; Kane, A.D.; Menon, D.K.; Glocker, B. Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med. Image Anal. 2017, 36, 61–78. [CrossRef]
D. Yi, M. Zhou, Z. Chen, and O. Gevaert, “3-D convolutional neural networks for glioblastoma segmentation,” pp. 1–8, Nov. 2016, [Online]. Available: http://arxiv.org/abs/1611.04534
O. H. Salman, M. F. A. Rasid, M. I. Saripan, and S. K. Subramaniam, “Multi-sources data fusion framework for remote triage prioritization in telehealth,” Journal of Medical Systems, vol. 38, no. 9, pp. 1–23, Sep. 2014, doi: 10.1007/s10916-014-0103-4.
S. M. R. K. Al-Jumur, S. W. Kareem, and R. Z. Yousif, “Predicting temperature of Erbil City applying deep learning and neural network,” Indonesian Journal of Electrical Engineering and Computer Science, vol. 22, no. 2, pp. 944–952, May 2021, doi: 10.11591/ijeecs.v22.i2.pp944-952.
S. Kareem and M. C. Okur, “Bayesian network structure learning using hybrid bee optimization and greedy search,” in 3rd International Mediterranean Science and Engineering Congress (IMSEC 2018), 2018, pp. 1–7. [Online]. Available: https://www.researchgate.net/profile/Shahab-Kareem/publication/333320417_Bayesian_network_structure_learning_using_hybrid_bee_optimization_and_greedy_search/links/5ce6aaf9458515712ebc96da/Bayesian-network-structure-learning-using-hybrid-bee-optimization-and-greedy-search.pdf
S. W. Kareem and M. C. Okur, “Pigeon inspired optimization of bayesian network structure learning and a comparative evaluation,” Journal of Cognitive Science, vol. 20, no. 4, pp. 535–552, Dec. 2019, doi: 10.17791/jcs.2019.20.4.535.
S. W. Kareem and M. C. Okur, “Bayesian network structure learning based on pigeon inspired optimization,” International Journal of Advanced Trends in Computer Science and Engineering, vol. 8, no. 1, pp. 131–137, 2019, doi: 10.30534/ijatcse/2019/2281.22019.
S. W. Kareem and M. C. Okur, “Evaluation of bayesian network structure learning using elephant swarm water search algorithm,” in Handbook of Research on Advancements of Swarm Intelligence Algorithms for Solving Real-World Problems, Hershey: IGI Global, 2020, pp. 139–159. doi: 10.4018/978-1-7998-3222-5.ch008.
Downloads
Published
How to Cite
Issue
Section
License

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.
All papers should be submitted electronically. All submitted manuscripts must be original work that is not under submission at another journal or under consideration for publication in another form, such as a monograph or chapter of a book. Authors of submitted papers are obligated not to submit their paper for publication elsewhere until an editorial decision is rendered on their submission. Further, authors of accepted papers are prohibited from publishing the results in other publications that appear before the paper is published in the Journal unless they receive approval for doing so from the Editor-In-Chief.
IJISAE open access articles are licensed under a Creative Commons Attribution-ShareAlike 4.0 International License. This license lets the audience to give appropriate credit, provide a link to the license, and indicate if changes were made and if they remix, transform, or build upon the material, they must distribute contributions under the same license as the original.





