Breast Cancer Screening Tool Using Gabor Filter-Based Ensemble Machine Learning Algorithms
Keywords:
Artefact, Cancer, Diagnostic, Gabor Filter, Radiologist, Random ForestAbstract
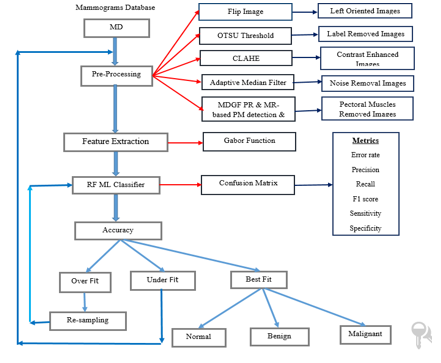
The most common kind of cancer among females that causes death is breast cancer. It's early detection and initial treatment can save the patient's life and also decrease the mortality rate. An efficient approach to finding breast cancer at an initial stage is screening mammography. However the diagnostic procedure is hand-operated, time-taking, and a specialist radiology is required and available only in hospitals, so the patient cannot check at their home with this technology. In the literature, many techniques have existed, but fail to produce a high accuracy rate due to the presence of noise, artefacts, pectoral muscles, and low contrast. Based on these reasons it is difficult for radiologists to find cancer at the initial stage. This paper presents the Gabor filter-based ensemble machine learning technique which gives a high accuracy rate in the presence of noise, artefacts, pectoral muscles, and low contrast. This method is applied on all MIAS Datasets, which consist of 322 mammogram images and produce an accuracy of 98.98%.
Downloads
References
Yan Wang , Zizhou Wang, Yangqin Feng , and Lei Zhang, “WDCCNet: Weighted Double-Classifier Constraint Neural Network for Mammographic Image Classification” IEEE Transactions on Medical Imaging, vol. 41, no. 3, March 2022.
Md. Akhlaqur Rahman, and Rajib Kumar Jha , “Multidirectional Gabor Filter-Based Approach for Pectoral Muscle Boundary Detection” IEEE Transactions on radiation and plasma medical sciences, vol. 6, no. 4, April 2022.
M. Broeders et al., “The impact of mammographic screening on breast cancer mortality in Europe: A review of observational studies,” J. Med. Screening, vol. 19, no. 1, pp. 14–25, Sep. 2012.
X. Shu, L. Zhang, Z. Wang, Q. Lv, and Z. Yi, “Deep neural networks with region-based pooling structures for mammographic imageclassification,” IEEE Trans. Med. Imag., vol. 39, no. 6, pp. 2246–2255, Jun. 2020.
L. Wang, L. Zhang, M. Zhu, X. Qi, and Z. Yi, “Automatic diagnosis for thyroid nodules in ultrasound images by deep neural networks,” Med. Image Anal., vol. 61, Apr. 2020, Art. no. 101665.
M. Mustra, M. Grgic, and R. M. Rangayyan, “Review of recent advances in segmentation of the breast boundary and the pectoral muscle in mammograms,”Med. Biol. Eng. Comput., vol. 54, no. 7, pp. 1003–1024, 2016.
M. A. Rahman, R. K. Jha, and A. K. Gupta, “Gabor phase response based scheme for accurate pectoral muscle boundary detection,” IET Image Process., vol. 13, no. 5, pp. 771–778, 2019.
R. Gupta and P. E. Undrill, “The use of texture analysis to delineate suspicious masses in mammography,” Phys. Med. Biol., vol. 40, no. 5, pp. 835–855, 1995.
P. K. Saha, J. K. Udupa, E. F. Conant, D. P. Chakraborty, and D. Sullivan, “Breast tissue density quantification via digitized mammograms,” IEEE Trans. Med. Imag., vol. 20, no. 8, pp. 792–803, Aug. 2001.
R. B. Gunderman, Essential Radiology. Clinical Presentation, Pathophysiology, Imaging, 2ed. New York, NY, USA: Georg Thieme 2006.
S. M. Kwok, R. Chandrasekhar, Y. Attikiouzel, and M. T. Rickard, “Automatic pectoral muscle segmentation on mediolateral oblique view mammograms,” IEEE Trans. Med. Imag., vol. 23, no. 9, pp. 1129–1140, Sep. 2004.
R. J. Ferrari, R. M. Rangayyan, J. E. L. Desautels, R. A. Borges, and A. F. Frere, “Automatic identification of the pectoral muscle in mammograms,” IEEE Trans. Med. Imag., vol. 23, no. 2, pp. 232–245, Feb. 2004.
V. B. Bora, A. G. Kothari, and A. G. Keskar, “Robust automatic pectoral muscle segmentation from mammograms using texture gradient and euclidean distance regression,” J. Digit. Imag., vol. 29, no. 1, pp. 115–125, 2016.
P. Shi, J. Zhong, A. Rampun, and H. Wang, “A hierarchical pipeline for breast boundary segmentation and calcification detection in mammograms,” Comput. Biol. Med., vol. 96, pp. 178–188, May 2018.
A. Rampun et al., “Breast pectoral muscle segmentation in mammograms using a modified holistically-nested edge detection network,”Med. Image Anal., vol. 57, pp. 1–17, Oct. 2019.
W. Cheung and G. Hamarneh, “N-SIFT: N-dimensional scale invariant feature transform for matching medical images,” in Proc. 4th IEEE Int. Symp. Biomed. Imag., Nano Macro, Apr. 2007, pp. 720–723.
R. Hupse and N. Karssemeijer, “Use of normal tissue context in computer-aided detection of masses in mammograms,” IEEE Trans. Med. Imag., vol. 28, no. 12, pp. 2033–2041, Aug. 2009.
G. Carneiro, J. Nascimento, and A. P. Bradley, “Unregistered multiview mammogram analysis with pre-trained deep learning models,” in Medical Image Computing and Computer-Assisted Intervention— MICCAI 2015 (Lecture Notes in Computer Science), vol. 9351. Cham, Switzerland: Springer, 2015, pp. 652–660.
K. Sankar and K. Nirmala, “An enhanced mammogram diagnosis using shift-invariant transform,” ICTACT J. Image Video Process., vol. 5, no. 2, pp. 920–925, Nov. 2014.
S. A. Taghanaki, Y. Liu, B. Miles, and G. Hamarneh, “Geometry-based pectoral muscle segmentation from MLO mammogram views,” IEEE Trans. Biomed. Eng., vol. 64, no. 11, pp. 2662–2671, Nov. 2017.
A. G. Gale, "The Mammographic Image Analysis Society digital mammogram database," Proceedings of the 2nd International Workshop on Digital Mammography, vol. 1069, pp. 375--378, 1994.
I. A. Inês C Moreira, Inês Domingues, António Cardoso, Maria JoãoCardoso, and J. S. Cardoso, "INbreast: toward a full-field digital mammographic database," Acad Radiol, vol. 19, no. 2, pp. 236-48, Feb 2012.
Sagenela Vijaya Kumar, C Nagaraju “Support vector neural network based fuzzy hybrid filter for impulse noise identification and removal from gray-scale image” Journal of King Saud University - Computer and Information Sciences, Volume 32, Issue 10, December 2020, Pages 1210-1211.
C Naga Raju, A Hima Bindhu “Primary Screening Technique for Detecting Breast Cancer” i-manager's Journal on Image Processing; Nagercoil Vol. 6, Iss. 2, (Apr/Jun 2019): 21-27.
Downloads
Published
How to Cite
Issue
Section
License

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.
All papers should be submitted electronically. All submitted manuscripts must be original work that is not under submission at another journal or under consideration for publication in another form, such as a monograph or chapter of a book. Authors of submitted papers are obligated not to submit their paper for publication elsewhere until an editorial decision is rendered on their submission. Further, authors of accepted papers are prohibited from publishing the results in other publications that appear before the paper is published in the Journal unless they receive approval for doing so from the Editor-In-Chief.
IJISAE open access articles are licensed under a Creative Commons Attribution-ShareAlike 4.0 International License. This license lets the audience to give appropriate credit, provide a link to the license, and indicate if changes were made and if they remix, transform, or build upon the material, they must distribute contributions under the same license as the original.





