Early Predictive Model for Detection of Plant Leaf Diseases Using MobileNetV2 Architecture
Keywords:
Deep Learning (DL), Support Vector Machines (SVM), Artificial Neural Networks (ANN), Mobilenetv2 model, K-Nearest Neighbour (KNN), Convolutional Neural Network (CNN)Abstract
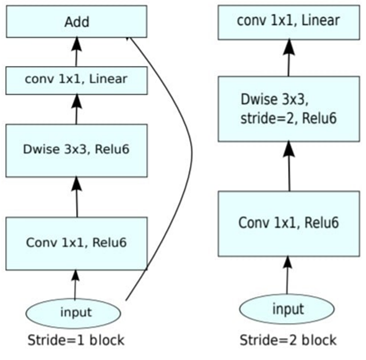
In farming, it is vital to recognize diseases of plant leaves and to improve the prediction quality of diseased plant leaves. Several laboratory-based techniques such as polymerase reaction decrease in agricultural output, and pesticide application, have really been identified for recognizing different diseases of plant leaves with human sight. Agriculture yield is improving every day as a result of current technological advancements. However, they are very time-intensive and costly for farmers. Deep Learning (DL) methods may help boost crop yields by identifying recently upgraded methodologies and diverse systematic patterns. To improve the reliability of the measurements, researchers focused on new methodologies in deep learning algorithms for diagnosing leaf diseases. Every model is essential and focuses on the path of deep learning applications as well as the challenges faced by farmers. The mobilenetv2 architecture is used in this study to determine how to diagnose diseases that affect leaves. This design is built on an inverted residual structure, with narrow bottleneck layers serving as the input and output of the residual block and extended representations as the inputs. Lightweight depth-wise convolutions are used in this architecture to select leaf characteristics in the intermediate expansion stage. This network has 53 layers in the very beginning. There are 32 filters in the first completely convolution layer, followed by 19 more bottleneck layers. To retain representational strength, non-linearities were eliminated in the narrow layers. The proposed method enables the decoupling of the input/output leaves from the transformation's expressiveness, offering a framework for additional study. On Imagenet classification, COCO object detection, and VOC picture segmentation, the performance is evaluated. The proposed model provides an accuracy of 95% in identifying the plant diseases.
Downloads
References
Madhulatha, G., and O. Ramadevi. "Recognition of Plant Diseases using Convolutional Neural Network." In 2020 Fourth International Conference on I-SMAC (IoT in Social, Mobile, Analytics and Cloud) (I-SMAC), pp. 738-743. IEEE, 2020.
D. Devarajan D Stalin Alex, T R Mahesh, Rajanikanth Aluvalu, V Vinoth Kumar, Uma Maheswari, S Shitharth, "Cervical Cancer Diagnosis Using Intelligent Living Behaviour of Artificial Jellyfish Optimized with Artificial Neural Network," in IEEE Access, 2022, https://doi.org/10.1109/ACCESS.2022.3221451
Rahman, Md Arifur, Md Mukitul Islam, GM ShahirMahdee, and MdWasiUlKabir. "Improved Segmentation Approach for Plant Disease Detection." In 2019 1st International Conference on Advances in Science, Engineering and Robotics Technology (ICASERT), pp. 1-5. IEEE, 2019.
Tetila, Everton Castelão, Bruno Brandoli Machado, Gabriel Kirsten Menezes, Adair da Silva Oliveira, Marco Alvarez, Willian ParaguassuAmorim, Nícolas Alessandro De Souza Belete, GercinaGonçalves da Silva, and HemersonPistori. "Automatic recognition of soybean leaf diseases using UAV images and deep convolutional neural networks." IEEE Geoscience and Remote Sensing Letters 17, no. 5 (2019): 903-907.
Ramakrishna, M.T.; Venkatesan, V.K.; Izonin, I.; Havryliuk, M.; Bhat, C.R. Homogeneous Adaboost Ensemble Machine Learning Algorithms with Reduced Entropy on Balanced Data. Entropy 2023, 25, 245. https://doi.org/ 10.3390/e25020245
Al Haque, ASM Farhan, Rubaiya Hafiz, MdAzizul Hakim, and GM Rasiqul Islam. "A Computer Vision System for Guava Disease Detection and Recommend Curative Solution Using Deep Learning Approach." In 2019 22nd International Conference on Computer and Information Technology (ICCIT), pp. 1-6. IEEE, 2019.
S. Roopashree, J. Anitha, T.R. Mahesh, V. Vinoth Kumar, Wattana Viriyasitavat, Amandeep Kaur, An IoT based authentication system for therapeutic herbs measured by local descriptors using machine learning approach, Measurement, Volume 200, 2022, 111484, ISSN 0263-2241, https://doi.org/10.1016/j.measurement.2022.111484
A. Ramcharan, K. Baranowski, P. McCloskey, B. Ahmed, J. Legg, and D. P. Hughes, “Deep learning for image-based cassava disease detection,” Frontiers in plant science, vol. 8, p. 1852, 2017.
P. Jiang, Y. Chen, B. Liu, D. He, and C. Liang, “Real-time detection of apple leaf diseases using deep learning approach based on improved convolutional neural networks,” IEEE Access, vol. 7, pp. 59069–59080, 2019.
Mahesh, T.R., Vinoth Kumar, V., Vivek, V. et al. Early predictive model for breast cancer classification using blended ensemble learning. Int J Syst Assur Eng Manag (2022). https://doi.org/10.1007/s13198-022-01696-0
S. Sladojevic, M. Arsenovic, A. Anderla, D. Culibrk, and D. Stefanovic, “Deep neural networks- based recognition of plant diseases by leaf image classification,” Computational intelligence and neuroscience, vol. 2016, 2016.
P. Chaitanya Reddy, R. M. S. Chandra, P. Vadiraj, M. Ayyappa Reddy, T. R. Mahesh and G. Sindhu Madhuri, "Detection of Plant Leaf-based Diseases Using Machine Learning Approach," 2021 IEEE International Conference on Computation System and Information Technology for Sustainable Solutions (CSITSS), 2021, pp. 1-4, doi: 10.1109/CSITSS54238.2021.9683020.
Mahesh, T. R., et al. "An Efficient Ensemble Method Using K-Fold Cross Validation for the Early Detection of Benign and Malignant Breast Cancer." International Journal of Integrated Engineering 14.7 (2022): 204-216.
M. Tahmooresi, A. Afshar, B. Bashari Rad, K. B. Nowshath, M. A. Bamiah, “Early detection of breast cancer using machine learning techniques,” Journal of Telecommunication, Electronic and Computer Engineering, vol. 10, no. 3-2, pp. 21-27, 2018.
Ebru Aydindag Bayrak, Pinar Kirci, TolgaEnsari,“Comparison of machine learning methods for breast cancer diagnosis.2019 Scientific Meeting on Electrical-Electronics & Biomedical Engineering and Computer Science (EBBT), pp. 1-3,2019.
Dursun Delen; Glenn Walker; Amit Kadam (2005). “Predicting breast cancer survivability: a comparison of three data mining methods”, Procedia Computer Science34(2), 113–127.
Asri, Hiba; Mousannif, Hajar; Moatassime, Hassan Al; Noel, Thomas (2016). Using Machine Learning Algorithms for Breast Cancer Risk Prediction and Diagnosis. Procedia Computer Science, 83(), 1064–1069.
Akbugday, Burak (2019). 2019 Medical Technologies Congress (TIPTEKNO) - Izmir, Turkey (2019.10.3-2019.10.5)] 2019 Medical Technologies Congress (TIPTEKNO) – “Classification of Breast Cancer Data Using Machine Learning Algorithms”,1–4.
T R, M., Kaladevi, A. C., J M, B., Vivek, V., Prabu, M., & Muthukumaran, V. (2022). An Efficient Ensemble Method Using K-Fold Cross Validation for the Early Detection of Benign and Malignant Breast Cancer. International Journal of Integrated Engineering, 14(7). https://doi.org/10.30880/ijie.2022.14.07.015 Retrieved from https://publisher.uthm.edu.my/ojs/index.php/ijie/article/view/10810
Qinghua Huang, Yongdong Chen, Longzhong Liu, Dacheng Tao, Xuelong Li, On combining biclustering mining and adaboost for breast tumor classification, IEEE Trans. Knowl. Data Eng. 32 (4) (2020)728–738.
Tamoor, Maria and Younas, Irfan. 'Automatic Segmentation of Medical Images Using a Novel Harris Hawk Optimization Method and an Active Contour Model'. 1 Jan. 2021: 721 – 739.
SapnaJuneja, AbhinavJuneja, GauravDhiman, SanchitBehl, SandeepKautish, "An Approach for Thoracic Syndrome Classification with Convolutional Neural Networks", Computational and Mathematical Methods in Medicine, vol. 2021, Article ID 3900254, 10 pages, 2021. https://doi.org/10.1155/2021/3900254.
K. K. Jha, R. Jha, A. K. Jha, M. A. M. Hassan, S. K. Yadav and T. Mahesh, "A Brief Comparison On Machine Learning Algorithms Based On Various Applications: A Comprehensive Survey," 2021 IEEE International Conference on Computation System and Information Technology for Sustainable Solutions (CSITSS), Bangalore, India, 2021, pp. 1-5, doi: 10.1109/CSITSS54238.2021.9683524.
Siddique, N.A., Paheding, S., Elkin, C.P., &Devabhaktuni, V.K. (2021). U-Net and Its Variants for Medical Image Segmentation: A Review of Theory and Applications. IEEE Access, 9, 82031-82057.
Dharahas, Reddy T., and T. R. Mahesh. "A Pragmatic Approach for Detecting Brain Tumors Using Machine Learning Algorithms." BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS Special Issue 14.11 (2021).
Mahesh, T. R., Vinoth Kumar, V., Muthukumaran, V., Shashikala, H. K., Swapna, B., & Guluwadi, S. (2022). Performance Analysis of XGBoost Ensemble Methods for Survivability with the Classification of Breast Cancer. Journal of Sensors.
R, M. T. ., Goswami, T. ., Sriramulu, S. ., Sharma, N. ., Kumari, A. ., & Khekare, G. . (2022). Cognitive Based Attention Deficit Hyperactivity Disorder Detection with Ability Assessment Using Auto Encoder Based Hidden Markov Model. International Journal of Communication Networks and Information Security (IJCNIS), 14(2), 53–65. https://doi.org/10.17762/ijcnis.v14i2.5464
Ramana, Kadiyala, Madapuri Rudra Kumar, K. Sreenivasulu, Thippa Reddy Gadekallu, Surbhi Bhatia, Parul Agarwal, and Sheikh Mohammad Idrees. "Early prediction of lung cancers using deep saliency capsule and pre-trained deep learning frameworks." Frontiers in oncology 12 (2022).
Downloads
Published
How to Cite
Issue
Section
License

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.
All papers should be submitted electronically. All submitted manuscripts must be original work that is not under submission at another journal or under consideration for publication in another form, such as a monograph or chapter of a book. Authors of submitted papers are obligated not to submit their paper for publication elsewhere until an editorial decision is rendered on their submission. Further, authors of accepted papers are prohibited from publishing the results in other publications that appear before the paper is published in the Journal unless they receive approval for doing so from the Editor-In-Chief.
IJISAE open access articles are licensed under a Creative Commons Attribution-ShareAlike 4.0 International License. This license lets the audience to give appropriate credit, provide a link to the license, and indicate if changes were made and if they remix, transform, or build upon the material, they must distribute contributions under the same license as the original.





